Let me share a frustration I never thought was possible: you look at a co-crystal structure of a protein with a small molecule bound to it and you have no idea what this small molecule is. That’s right. I have to admit that this is a fairly awkward feeling for someone who has a lab that makes small molecules. My students rely on X-ray crystallography for structure determination in our day-to-day endeavours. However, we use direct methods in organic chemistry and, sure enough, when you have a molecule whose structure is solved at 0.6 Angstrom resolution, life is good and you can rely 100% on the data when it tells you what a given molecule is. Protein structures are way more complex, yet rely on a MODEL when you try to solve the structure. If you have a 2.4 Angstrom resolution structure and you see electron density corresponding to a small molecule bound to the protein, good luck finding out what this molecule is unless you already know it or suspect what it might be! Isn’t it frustrating?
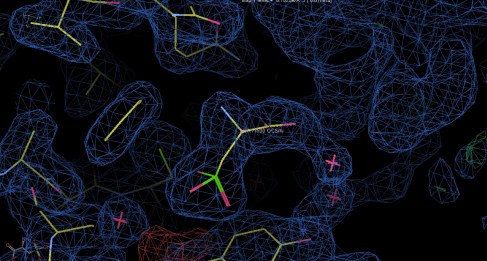
A case in point is shown below, which is our day to day struggle at the moment… Right in the center of the picture you see a banana-like electron density which is not part of the protein. You do see a (presumed) structure and it is shown as a model. But don’t be fooled, this is far from reality. We know that it ain’t real since anomalous scattering tells us that there are no S atoms, despite the SO3- group clearly modelled in the structure shown. We are trying to find out what the density belongs to and are still in the dark. I never thought (perhaps naively) that this could be the case, but it is true… Protein crystallography is for proteins and small molecules appear to be the proteins’ ugly cousins! We are trying to use all sorts of tools, including non-denaturing MS but we can’t seem to zero in on the right structure (yet). Stay tuned. Incidentally, this is a protein I crystallized with Elena not long ago. I can’t disclose the name of it yet, sorry.
I love this sabbatical!