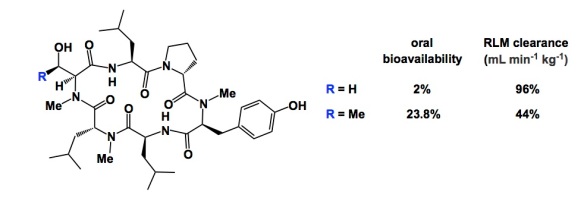
Over the past several days, I have been suspiciously absent from the blogosphere. I was on a lecture trip to California, during which I visited the University of California, Santa Cruz and the University of California, San Francisco. The usual thing: you arrive in the morning, you give your talk, and you get exposed to the latest developments in other labs. Nothing can beat this way to spend time. I am very grateful to my hosts – Professor Scott Lokey in Santa Cruz (http://www.chemistry.ucsc.edu/faculty/singleton.php?&singleton=true&cruz_id=slokey) and Professor Jack Taunton (http://cmp.ucsf.edu/faculty/jack-taunton/) in San Francisco. Tonight I will talk about Scott. He and I share a common passion for macrocycles. I think you are all familiar with the byzantine difficulties in forcing cyclic peptide molecules to “behave”. I refer to the chasm that exists between small molecules and cyclic peptides when it comes to drug-like properties. Scott’s lab is well known for its findings that have been reverberating through the scientific community. In particular, I refer to his ongoing research that shows how important hydrogen bonds are in maintaining the conformations of complex macrocycles. Here is a cool example from the Lokey lab that tells you that we are still far from understanding this class of compounds. What you see is that the serine derivative has 96 mL min-1 kg-1 RLM clearance (RLM: rat liver microsomes), whereas the threonine-containing congener is substantially less stable (44 mL min-1 kg-1 RLM clearance). Oddly enough, the serine-containing peptide actually has an oral bioavailability of only 2% compared to the threonine-containing peptide which is 23.8% orally bioavailable. The difference between these two molecules is just one methyl group…
Click to access c2md20203d.pdf
This example underscores the highly empirical nature of efforts to identify orally bioavailable macrocycles and suggests that finding a correlation between oral bioavailability and scaffold design is likely to be challenging. I think it will take us all a long time to understand the intricate factors that turn cyclic peptides into drug-like molecules. I have no doubt that it will eventually happen and, when it does happen, we will probably collectively quote Winston Churchill, who famously said: “We always come to the right decision, having tried everything else first”.