Here it is, guys. The structure is 2.4 Angstrom resolution, which is not too bad for a start. This happens to be a PWWP domain that recognizes methylated lysine containing peptides. The goal now is to design ligands on the basis of the small molecule fragment (you can see it) that has co-crystallized. I was talking to my PhD student Rebecca the other day and she is very keen to tackle this problem. Today I learned that another crystal might be sent out to the synchrotron soon. That will be awesome. Big-time thanks to Elena (who teaches me protein crystallography) and to Aiping Dong, who is the crystallography expert. He is amazing. I love chasing proteins!
Monthly Archives: July 2013
Boston, Novartis, and good times
I flew into Boston last night to give a talk at Novartis. I had a great time there today. Donovan Chin (Senior Investigator at the NIBR) was my host. I had a lot of fun hearing about their interests in molecules that are also dear to our hearts. I am talking about peptide macrocycles. Scott Lokey of the UCSC was also here at Novartis last week. Scott is doing some trailblazing work trying to understand the cell permeability of cyclic peptides.
It is clear that the industry appetite to macrocycles is high, particularly with regard to gauging their complex stereochemical preferences… It is eye-opening to talk to people in industry who do this for a living. When I hear that docking a small molecule takes a couple of minutes (I refer to computational docking into a binding site of a protein target), whereas the same process for a macrocycle is close to 18 hours, I really wonder how anyone will get the “heavy lifting” (to use the language of Dr. Andrew Roughton, the COO of Encycle Therapeutics, a company I founded in 2012) done in this space. It is, nonetheless, a fascinating problem and we are going to deal with it heads-on as a community. Naively, I asked Donovan about why does it really matter to be “right” in a complicated maze of conformations that are differentiated by a fraction of a kcal/mol. However, it is important to keep in mind that this thing is additive… To use Donovan’s analogy, it is like walking in the streets of Paris (or any other city with a complex web of streets): one wrong turn and your mistake is exacerbated such that the “real destination” will never be found… If we have a cycle that is composed of (say) 8 amino acids – once you add all the rotors, it is easy to see that this is a tough problem to crack computationally.
We needed to have some beers at the end of this day and went to this place called “Catalyst” right next to MIT. My former student Tim Rasmusson, who works at Novartis, came out – it was cool to see him! He is doing great. I hoped Ben Rotstein and Zhi He might join us (former PhD students who are now doing their PDFs at Harvard and MIT, respectively), but they were busy. I am at the airport now and will blog about my protein crystals tomorrow. The data is in!
Not making covalent bonds
So… I have a ton of work this weekend and it is mainly about making constructs for our next protein expression experiments. I gotta tell you: this is opening my eyes to entirely new challenges. See, I have been trained as a synthetic organic chemist, which means (more or less) that we think how to forge COVALENT bonds between atoms. Seriously, this is what we do in synthesis (in rough terms). Typically, good things that are new and interesting come from irreversible connections between atoms (in a way protein expression is still synthesis, but you think on a different level that does not involve worrying about how the bonds are constructed). I now must admit that there are really interesting challenges out there I have not considered seriously until now. The biggest difference from what I have been trained to do is that covalent bonds do not drive progress anymore. For example, this weekend I need to think of constructs for several receptor proteins that we will hopefully be able to express in E. coli. This is a different ball game altogether and it has its own unique challenges. Not only do we want to make clones but we also want to make sure we will be able to express the proteins. The ultimate goals are varied and range from crystallization to the production of antibodies. However, I catch myself thinking that: a. I am excited about what I do; b. it is new to me and c. whatever I do is NOT about carefully orchestrating covalent bond making. This latter point is important because I have never thought that I would be interested in science that does not, ultimately, result in making new bonds. But here I am – making constructs for expression and I do not think about gluing things together. Kind of different and really interesting!
Proton/deuterium exchange
There are no protein reports today… I am waiting to hear on the news from the synchrotron (tomorrow). For a change, today I am going back to MUCH smaller molecules. I was thinking if I can come up with an interesting question involving some really well known reagents that seem almost too trivial. The venerable borohydride came to mind! Take a look at the equilibrium shown below. This process can be driven to the right, offering a means to make borodeuteride. The finding comes from a rather old paper (I can post it in the future). I did not balance the equation and decided to just leave the bare bones of the central idea. What do you think is the mechanism of this exchange process? 
On protein synthesis, ligation, and expression…
Some of you know me – I love ligation technologies. As a chemist, it is exciting that synthesis can deliver a protein. Cool stuff. Now… Here comes a question I would not have asked had I not tried protein expression myself. In one of our recent experiments, Elena and I got about half a gram of a particular protein (more on that in the future) from E.coli in about 3 days. This technology is really unbeatable when it works. You don’t need to worry about epimerization (the way we do when ligations are considered), folding, etc. I am just curious – out of scientific publications that describe protein synthesis using modern chemical ligation methods, how many actually describe syntheses that CANNOT be performed using expression methods? Because — I tell you what (having done this myself) — you just cannot beat expression. OK, I understand that not all proteins express well and this is a good argument… But still – are you telling me that whenever people publish a chemical synthesis of a protein type of paper, they preamble their work by saying that this synthesis CANNOT be achieved using expression methods? If such synthesis can be done using expression, I honestly just do not see why you would use any method other than expression…
P.S. You want post-translationally modified proteins? Use insect or mammalian expression. From what I saw, folks at SGC do that routinely…
First protein crystals
Today we shipped a bunch of our newly prepared protein crystals to be analyzed on the synchrotron. The process of getting crystals is not exactly science… It is a form of art and I am in awe of all those people who have contributed (over the years) to streamlining this process. Below I am showing one of the results of today’s work. The size of the well is about 2.5mm, which makes the crystal about 0.5mm (I guess… Give or take…). We got this bad boy shipped out, the results should be back on Friday. This, by the way, is by no means the final product!
In fact, we needed to painstakingly fish out this crystal with a micro-lasso sort of a tool (below), suspend it in special oil to get rid of water, and rapidly freeze it in liquid nitrogen (all the while trying to avoid the crystal from forming cracks).

You can see the holder with mounted crystals immersed in liquid nitrogen in the photo below.
The beast diffracted on the SGC machine but we’ll see the news from the synchrotron beam!
Of course I did not say what this protein is… Let me just say this: it is one of the trimethyllysine binding domains. I will elaborate on this in the future (and you will hopefully see where we are planning to take this from here).
My crystalline sabbatical
Hey guys, I mentioned that I am on sabbatical from July till the end of December, 2013. I will elaborate. For a number of reasons I chose not to leave Toronto but to stay here and concentrate on something that is totally new to me. With this in mind, I joined SGC (Al Edwards, its director, is an old friend of mine). Here is their website:
http://www.thesgc.org/scientists/groups/toronto
I now work with Elena Dobrovetsky. She is awesome. I am learning a ton about protein crystallography, all the way from expression to mounting of crystals and data collection. I will be posting my progress reports. Just 2 weeks ago we set up 960 crystallizations of some epigenetic domains (more on that later) and got some really nice crystals already (actually only one condition delivered – attrition was high). My ultimate goal is fragment based screening using crystallography. This stuff is a ton of fun. I can hardly be found at my main office (Lash Miller Building), but I am fine with that – if you do a sabbatical, you may as well try to see that there really is a separation from your regular job. I asked my wife “Do you think my students miss me?” and she said “Ha ha ha! No way!!”. I suppose this is true… I am not sad though!
Boron in peptides
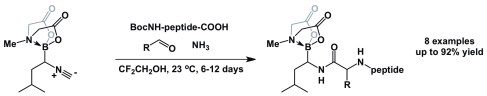
Here is our most recent addition to the lineup of amphoteric reagents. I refer to alpha-boryl isocyanides. These molecules enable synthesis of boron-containing peptides and heterocycles. Adam Zajdlik (about to start his second year of PhD in my lab) developed these species… Check out the photo – these are white solids (and they do not stink the way most isocyanides do… We call them “peer-friendly” isocyanides…)

http://onlinelibrary.wiley.com/doi/10.1002/anie.201302818/abstract
If you want to know some history, the isocyanides emerged on the heels of some superb foundational work by my former PhD student Zhi He (now doing his post-doctoral work with Tim Jamison at MIT). Zhi developed alpha-boryl aldehydes using MIDA group on boron:
http://pubs.acs.org/doi/abs/10.1021/ja205910d
We were happy to publish this chemistry in response to questions I have been getting at conferences ever since we disclosed the aziridine aldehyde synthesis with Ryan Hili. People were asking me: “OK… the amine/aldehyde combo is cool, but can you think of a molecule where a carbon nucleophile is orthogonal to the aldehyde group?” Thanks to Zhi, we have an answer!
I want to end this post by saying that the unusual stability of our growing line-up of borylated reagents is due to the MIDA functionality, which has been popularized by Marty Burke (http://www.scs.illinois.edu/burke/). Marty has done some pioneering work with MIDA boronates.
What is “amphoteros”
The word amphoteros (the name of this blog) comes from Greek and means “both worlds”. We use this term to describe molecules that have two opposing properties. You might think such molecules should be unstable, but we know how to force them to “behave”. Here is one we developed with Ryan Hili (now at the University of Georgia):
http://pubs.acs.org/doi/abs/10.1021/ja065898s
Ever since this 2006 paper, my lab has been busy exploring the strange world of amphoteric molecules… One of the things you will read in my blogs is how we go about developing this chemistry. You will also learn about the students who have put their hard work into this area.
It’s live!
After years of contemplating science blogging, I decided to roll up my sleeves and get going… I think this is where science publishing (at least some of it) is likely heading. This is my main motivation. As well, I just want to let people know about what we do in my lab at the University of Toronto:
http://www.chem.utoronto.ca/wp/yudinlab/
Now that I am on sabbatical until the end of December 2013, I have some more time – so here you go!