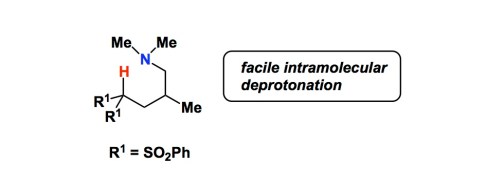
Synthetic organic chemists have always been awestruck by the remarkable rate enhancements on display in enzymatic reactions. One can list many different attributes possessed by nature’s catalysts. I want to talk about the most important one (in my view): control over solvation/desolvation. Is there anything synthetic that has this property? Menger has introduced the concept of spatiotemporal control, asserting that intramolecular reactions occur at enzyme-like rates when van der Waals contact distances between the reacting groups are too small to accommodate intervening solvent. For example, if two reactive groups are held in such a way that a water molecule can position itself between them, the groups are not considered to be in proximity. If we assume that water molecule is about 3Å in diameter, distances less than 3Å are expected to result in superfast intramolecular reactions. Here is an example. Enzymes are known to remove protons from fairly non-acidic CH groups using weakly basic histidine side chains (pKa of imidazole = 7). A synthetic model shown below has the CH-N distance of 2.34Å (which is less that 2.75Å sum of van der Waals radii of H and N). As a result, a fast intramolecular general-base-catalyzed proton exchange between the amino group and the methine proton takes place (measured through H/D exchange). This exchange is too fast to measure by NMR at -80 oC! In contrast, the intermolecular control is too slow to measure at 100 oC… The pKa’s of the amine conjugate acid and the CH bond marked below are 10 and 13.8, respectively. This example provides a “sneak peek” at what happens within hydrophobic pockets of enzyme active sites. Needless to say, synthetic chemists are keen on events that unfold on scales far greater than 3Å, which is why the so-called “enzyme models” never worked, do not work, and will never work. There’s just too much solvent around, once our overengineered molecules with all those cleverly crafted side chains are subjected to scrutiny. They might be decent catalysts, but they ain’t enzymes. My postdoctoral advisor, Barry Sharpless, once gave a talk entitled “The Secret Lives of Enzymes”. I think there is no secret, except for the unsurpassed control of solvation and desolvation.
Monthly Archives: November 2013
Structure chasers
I am going to talk about the work done by Serge Zaretsky and Dr. Conor Scully in my lab. Serge is a PhD student, whereas Conor is a research associate. One of our long-standing goals is to understand the structure/function relationship of complex macrocyclic structures. We are not the only ones who do this sort of research as there is a real gold rush to identify peptide macrocycles that approach the notorious cyclosporine molecule in terms of cellular permeability. This, of course, is just one of the many interesting properties macrocycles might possess. I think what distinguishes our work is a firm commitment to detailed structural studies, which is not something people do all the time, especially in the peptide community. I think this interest in structural characterization stems from our work in small molecule chemistry, which has been our main methodological focus.
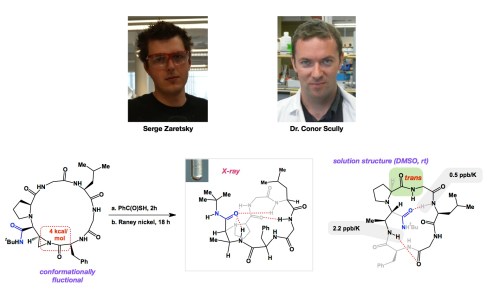
Serge and Conor have developed a robust system that enables us to quickly identify a range of control elements in macrocycles. In this regard, I would be remiss if I did not mention the pioneering efforts of Dr. Vishal Rai (currently a faculty member at IIT Bhopal, India). Vishal did his postdoctoral work in our lab several years ago. In terms of our structural work, we endeavour to understand complex macrocycles using a combination of NMR and X-ray crystallography. Below is a graphic and a link to Serge and Conor’s recent article. An aziridine-containing macrocycle is a typical entry point into our efforts. Because aziridine nitrogen does not conjugate well with the nearby carbonyl group, this particular structure is very fluxional. However, when you engage the three membered ring through nucleophilic ring-opening, things change. The structure folds in such a way that the side chain (in blue) acts as a centerpiece, organizing the hydrogen bonds around it. Importantly, this situation holds in solution, yet in a different way. The third graphic corresponds to our recently determined solution NMR structure and shows how the blue group “swings” underneath the ring compared to the position it holds in the crystalline state. The numerical values in grey stand for chemical shift changes as the temperature is changed. This is a good way to chase hydrogens that are involved in hydrogen bonds.
http://onlinelibrary.wiley.com/doi/10.1002/chem.201303453/suppinfo
There are many lessons one can extract from this study. One possibility is to use the “blue bond” behavior and thread side chains through the center of the ring, which has always been a dream of mine. I think that Mother Nature is trying to tell us something by showing how the blue group is “glued” to the center of the ring in both the solid and solution states alike. We will probably need to spend some time optimizing the size of the macrocycle for our “threading” purposes. I suppose we might learn a lesson from the following natural product structure described by Nar and co-workers (pdb code 3NJW). You will note a fairly small (by protein standards) ring and a magenta thread woven through the middle. Incidentally, this is the first X-ray structure of a lasso peptide! This beautiful structure is exceptionally stable against proteolysis.
http://onlinelibrary.wiley.com/doi/10.1002/cmdc.201000264/abstract
Electrostatic vs orbital control
Electrostatic vs orbital control has always been challenging to teach, at least for me. Indeed, the boundaries are sometimes blurred. We all know what each of these concepts mean. Electrostatic interactions are due to the presence of ionized chemical entities and to the electronegative and electropositive properties of atoms. On the other hand, orbital interactions are between frontier molecular orbitals that belong to the reaction partners and need not be polarized (e.g. a Diels-Alder reaction between butadiene and ethylene). Earlier today, I had to revisit some classic papers by Professor Tanner of Denmark. This work describes his lab’s efforts towards the total synthesis of balanol, which is a neat natural product that has activity (not amazing by today’s standards) against protein kinase C. While reading this work, I got reminded about a really nice example of electrostatic vs orbital control. In the Tanner case, it is electrostatics that rule. Shown below are two cases – an aziridine and epoxide opening. Each example leads to ring-opening with very high regioselectivity. You will note that C4 position is being hit preferentially in both cases, despite the obvious similarity between C3 and C4 surroundings. There may be some steric effects that govern this selectivity, but the authors ascribe their findings to electrostatic control. According to Tanner’s calculations, there is very little LUMO component on each of the methines in the three-membered rings. However, there is a notable difference between partial charges and they correlate with regioselectivity. If it were orbital control – how can one possibly imagine that fluoride and azide would give the same major regioisomer (they also did cyanide and cuprate – all go C4!)? In my view, this is a good example of electrostatic control. I would add, though, that contemporary computational approaches may provide additional clarification of this phenomenon.
http://www.sciencedirect.com/science/article/pii/S0040402097001671
Neighbourly ties
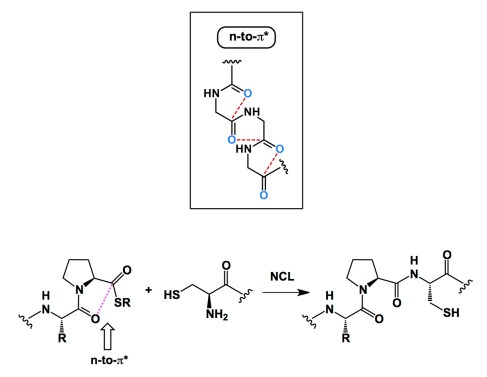
In 2009, Raines and Woolfson published their influential paper on the concept of n-to-pi* interaction in proteins (http://www.nature.com/nchembio/journal/v6/n8/full/nchembio.406.html). By analyzing the protein databank they came to the conclusion that there exists a previously unappreciated attractive interaction between backbone amides. This interaction arises from the delocalization of a lone pair of electrons (n) from an oxygen atom to the antibonding orbital (pi*) of the subsequent carbonyl group. For synthetic organic chemists this sort of interaction makes sense. Who can forget the Burgi-Dunitz trajectory of attack? However, it has gone under the radar of structural biologists, which is somewhat surprising (or not). Given the significance of this interaction, there is a push to adjust parametrization of many molecular modeling packages. There are many other implications, but what about peptide reactivity? I can think of one cool example. It comes from Kent and co-workers, who published an interesting Chem Comm paper not too long ago (http://pubs.rsc.org/en/content/articlehtml/2011/cc/c0cc04120c). In it, the authors explain the relatively low reactivity of prolyl thioesters in native chemical ligation. According to Kent, the n-to-pi* interaction is playing a role in diminishing the electrophilicity of the thioester.
Building blocks for peptides
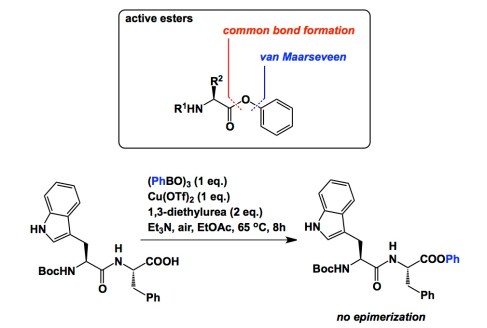
There are many ways to make a peptide bond, which is one of the most ubiquitous linkages known to man. Regardless of the biosynthetic or synthetic origins of a particular process, the acid component is turned into a better electrophile during peptide bond formation. DCC couplings and active ester technologies do this job for countless chemists around the world. If you look at the active ester method, you will note that there is one major way of producing active esters: using the red disconnection marked below. But what about an alternative pathway shown in blue? This is a viable option, yet no one has thought of it prior to Jan van Maarseveen’s recent work in Chem. Eur. J. I talked with Jan about this project during my trip to the Netherlands this past June. The reaction is an “outside the box” way of looking at problems, which prompts “Why didn’t I think about this?” kind of question. Really, when you first consider this idea, you expect that it must have been tried prior to 2013 by the legions of synthetic chemists eager to test new substrates in coupling reactions. As it turns out, Jan was the first one to think about this application and reduce it to practice. My complements. Of note is the configurational stability of the alpha-stereocenter during the coupling reaction. This chemistry should open up many new synthetic possibilities.
http://onlinelibrary.wiley.com/doi/10.1002/chem.201303347/abstract
Lemons, lemonade, and bromosporine
I have been in the UK for the past couple of days, visiting the SGC site at Oxford University. Paul Brennan has been my host here (http://www.ndm.ox.ac.uk/principal-investigators/researcher/paul-brennan). I thoroughly enjoyed my visit because I have got to interact with Paul and his group as well as a number of other PIs at the SGC. Paul showed me some of his recent science pertaining to the development of bromodomain inhibitors. Bromodomains are protein interaction modules that recognize ε-N-lysine acetylation motifs. I already mentioned them when I talked about isoxazoles. By the way, the name “bromodomain” has nothing to do with bromine. There are no bromine atoms in this protein, or nowhere close for that matter. Instead, the name appears to be related to the brahma (brm) gene, which is required for the activation of multiple homeotic genes in Drosophila. Don’t ask me how “brahma” became “bromo” as I do not think anyone has a clue. In terms of bromodomain inhibitors, there is an interesting story that has developed here in Oxford over the past year or so. Paul’s group has designed the so-called bromosporine (another interesting name… think about cyclosporine and staurosporine…). You can read about it at the following link: http://www.thesgc.org/scientists/chemical_probes/bromosporine. The strength of this molecule is in its promiscuity, which offers an interesting lesson to all of us. Promiscuous inhibitors such as bromosporine are invaluable in assay development due to their broad-spectrum mode of action. Indeed, when you have lemons – you make lemonade!
Bromosporine is a potent inhibitor of a range of bromodomain proteins. Consistent with SGC’s mantra, bromosporine has been released to the public and can now be purchased from a number of commercial sources. When I saw Paul’s nice synthesis of this molecule, it reminded me about the virtues of nucleophilic aromatic substitution. In the old days, pyridiazine was the workhorse of many ligand syntheses in the Sharpless lab. In Paul’s case, substituted trichloropyridazine is the starting electrophile that is subjected to treatment with hydrazine, which is followed by heterocycle formation using acetic acid. It is instructive to ponder about the regioselectivity of the first and second steps (which gives me an idea for a good cumulative exam question). Incidentally, it is the 5-membered heterocycle unit that acts as the acetyllysine mimic. This is seen in the co-crystal structure, although I do not think it has been deposited into the pdb thus far.
I am now off to London for a day. I’ve heard Natural History Museum has an interesting Darwin exhibit.
Empowering students with strong synthetic skills
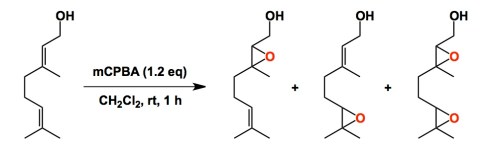
From a sheer pedagogical perspective, one of my all-time favorite papers is the well-crafted 2-page J. Chem. Ed. article by Tom Hoye and co-workers from the University of Minnesota. The goal of J. Chem. Ed. is to publish papers that serve as useful teaching tools. Hoye’s paper concerns with the epoxidation of geraniol. Sounds easy? The caveat is that the students have to perform this chemistry using 1mg of the starting material. The three epoxides derived from geraniol differ substantially in terms of their Rf. The students have to isolate and identify each product using conventional chromatography techniques. Each pure fraction is supposed to be characterized by 1H NMR. I know that the Chemistry Department at the University of Minnesota used this exercise as part of their teaching curriculum.
http://pubs.acs.org/doi/abs/10.1021/ed083p919
Organic synthesis is an experimental science and such small-scale experience is invaluable. According to Tom, less than 1% of new graduate students can do this with confidence. So, if you are a beginning (or an intermediate) synthetic chemist – my suggestion is that you give this reaction a shot in your spare time (using a 1 mg scale!). You do not need to tell the result to your advisor, but you will know if you need to put more work into developing better command of experimental techniques. According to Tom, the ability to perform in this test correlates really well with experimental acumen across the entire range of preparative techniques.
Psychophysical sensations
The province of Szechuan, China, is widely regarded for its culinary delights. I applaud the Szechuan people’s commitment to food that feels suicidally hot. On the molecular level, the so-called Szechuan pepper is behind those dishes. The closest we get to this kind of food in Toronto is through a couple of restaurants in Chinatown that are notorious for their spicy Szechuan cuisine. Here is a shot of my lab at a group outing last week. I am also showing you alpha-hydroxy sanshool, the culprit present in those infamous Szechuan peppercorns. These peppercorns look weird, nothing like your typical pepper. Unfortunately, not everyone was able to join us in this journey (Jeff was running a TLC, whereas Serge probably wasn’t brave enough…).
Now I understand why my tongue is numb every time I have Szechuan food. It is because alpha-hydroxy sanshool excites my neurons by inhibiting several (yes, not just one!) anesthetic-sensitive potassium ion channels. Here is a Nature Neuroscience paper that goes into the “unique and complex psychophysical sensations associated with the Szechuan pepper experience”.
http://www.nature.com/neuro/journal/v11/n7/full/nn.2143.html
Shifting sigma bonds in concert
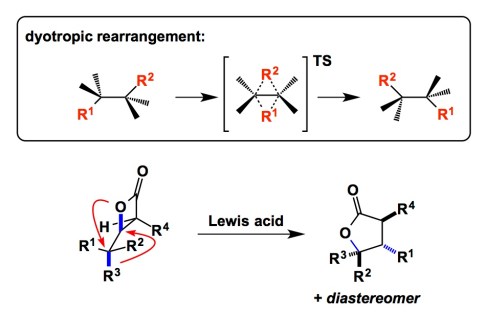
Nowadays, when one talks about discovering new reactions, one quickly realizes that there’s not a whole lot of fundamentally new elementary processes that remain unknown. As a result, novel reactions tend to be “composed” of different permutations of well-known elementary processes. Every now and then a mechanistically distinct transformation pops up. I keep an eye on reactions of that sort. Daniel Romo’s elegant experimental work coupled with Dean Tantillo’s theoretical approach provide a glimpse at some useful, yet fairly uncommon, types of reactions – the so-called dyotropic processes (see the graphic below). In sigmatropic rearrangements, a pi/sigma-system undergoes a transformation that results in a net translocation of one sigma bond and concomitant shift of the pi-system. In contrast, dyotropic rearrangements describe simultaneous migration of two sigma-bonds. Evidence in the JACS report cited below suggests that, depending on the nature of the Lewis acid, a concerted or stepwise mechanism takes hold. While reactions of this kind have been known for some time, they have not reached the mainstream of synthesis. I think there is a lot of room for reaction discovery using this mode of reactivity in the context of complex electrophiles.
http://pubs.acs.org/stoken/beta/select/abs/10.1021/ja303414a
Displacement of ice rings
Small molecules come in all sorts of different shapes and are capable of wonderful things. Of particular significance are those privileged types of small molecules that interact with their protein targets with high ligand efficiency. I will dedicate a future blog post to the concept of ligand efficiency, but I am sure that we all “feel” what it means without any formal definition. Ligand efficiency is a measure of how well a given molecule engages its innate features when it interacts with a protein target. Understanding the balance of enthalpy and entropy in this type of molecular recognition is paramount in probe/drug design. Great strides have been made in explaining (through docking simulations) how and why organic molecules interact with their targets. A lot of the developed algorithms work extremely well (Schrodinger’s Glide package is my favorite at the moment). The simulations may not have the best predictive power, but they are certainly capable of explaining experimental facts. Or are they?
I give you one of the workhorses of molecular biology: biotin/streptavidin interaction. Everyone is aware of the significance of this non-covalent “glue”. There are many examples of affinity experiments that utilize the strength of biotin/streptavidin pair. Personally, I have been in awe of this system because I just don’t get it: how the heck is such tight binding possible? You look at the dinky little biotin molecule and there is just no way to expect that its binding interaction with streptavidin would be on the order of 10-14 mol/L (I mean the dissociation constant, Kd). It turns out that sophisticated modern simulations have a heck of a trouble predicting this extreme ligand efficiency. The experimentally determined binding affinity is ORDERS of magnitude higher than predicted. So what is the basis for this impressive affinity?
It turns out that the reason likely lies in the amazing five-membered, ice-like ring of water molecules that is present in the binding site of streptavidin. Biotin is uniquely geared towards displacement of this “ice-ring” arrangement from the binding site, which generates a huge entropic driving force (see the graphic above showing the ring and the ultimate structure where biotin is in its place). There is still no way to predict this kind of behaviour computationally, I am afraid. The lesson here is that displacement of ordered waters is a VERY attractive way towards designing super-ligands. Berne and co-workers have published a very insightful PNAS paper on this subject several years ago (I thank Robert Campbell of Queens University for bringing this work to my attention):